pyribs¶
Discord | Mailing List | |
|---|---|---|
Website | Source | Docs | Paper |
|---|---|---|---|
PyPI | Conda | CI/CD | Docs Status |
|---|---|---|---|
A bare-bones Python library for quality diversity (QD) optimization. Pyribs implements the highly modular Rapid Illumination of Behavior Space (RIBS) framework for QD optimization. Pyribs is also the official implementation of Covariance Matrix Adaptation MAP-Elites (CMA-ME), Covariance Matrix Adaptation MAP-Elites via a Gradient Arborescence (CMA-MEGA), Covariance Matrix Adaptation MAP-Annealing (CMA-MAE), and scalable variants of CMA-MAE.
Community¶
Join the Pyribs Announcements mailing list for infrequent updates (less than 1/month) on the status of the project and new releases.
Need some help? Want to ask if pyribs is right for your project? Have a question that is not quite a bug and not quite a feature request?
Join the community Discord!
Overview¶
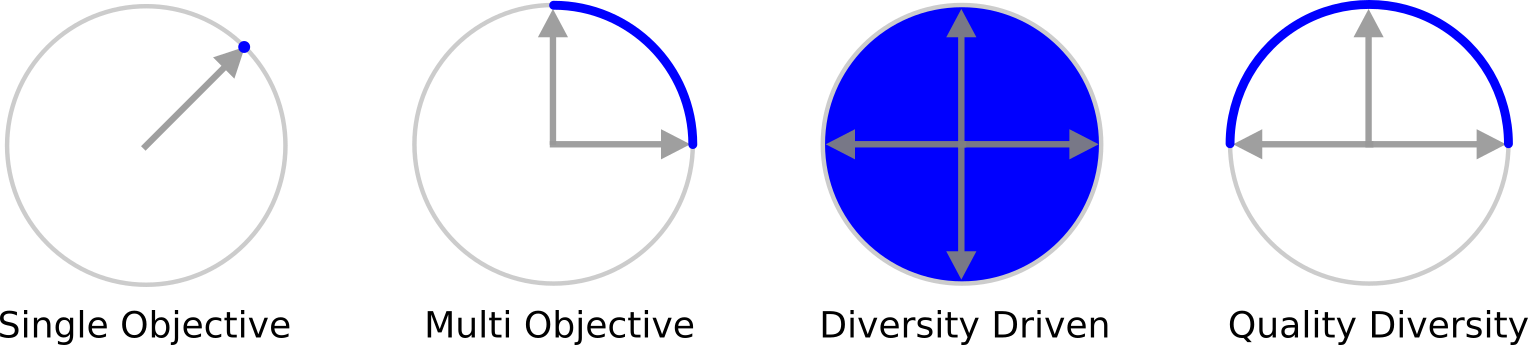
Quality diversity (QD) optimization is a subfield of optimization where solutions generated cover every point in a measure space while simultaneously maximizing (or minimizing) a single objective. QD algorithms within the MAP-Elites family of QD algorithms produce heatmaps (archives) as output where each cell contains the best discovered representative of a region in measure space.
In the QD literature, measure function outputs have also been referred to as “behavioral characteristics,” “behavior descriptors,” or “feature descriptors.”
Recent years have seen the development of a large number of QD algorithms. To represent these and future algorithms, we have developed the highly modular RIBS framework. RIBS divides a QD algorithm into three components:
An archive, which saves generated solutions within measure space.
One or more emitters, where each emitter is an algorithm which generates new candidate solutions and responds to feedback about how the solutions were evaluated and how they were inserted into the archive.
A scheduler that controls the interaction of the archive and emitters. The scheduler also provides an interface for requesting new candidate solutions and telling the algorithm how candidates performed.
By interchanging these components, a user can compose a large number of QD algorithms.
Pyribs is an implementation of the RIBS framework designed to support a wide range of users, from beginners entering the field to experienced researchers seeking to develop new algorithms. Pyribs achieves these goals by embodying three principles:
Simple: Centered only on components that are absolutely necessary to run a QD algorithm, allowing users to combine the framework with other software frameworks.
Flexible: Capable of representing a wide range of current and future QD algorithms, allowing users to easily create or modify components.
Accessible: Easy to install and learn, particularly for beginners with limited computational resources.
In contrast to other QD libraries, pyribs is “bare-bones.” For example, like pycma, pyribs focuses solely on optimizing fixed-dimensional continuous domains. Focusing on this one commonly-occurring problem allows us to optimize the library for performance as well as ease of use. Refer to the list of additional QD libraries below if you need to handle additional use cases.
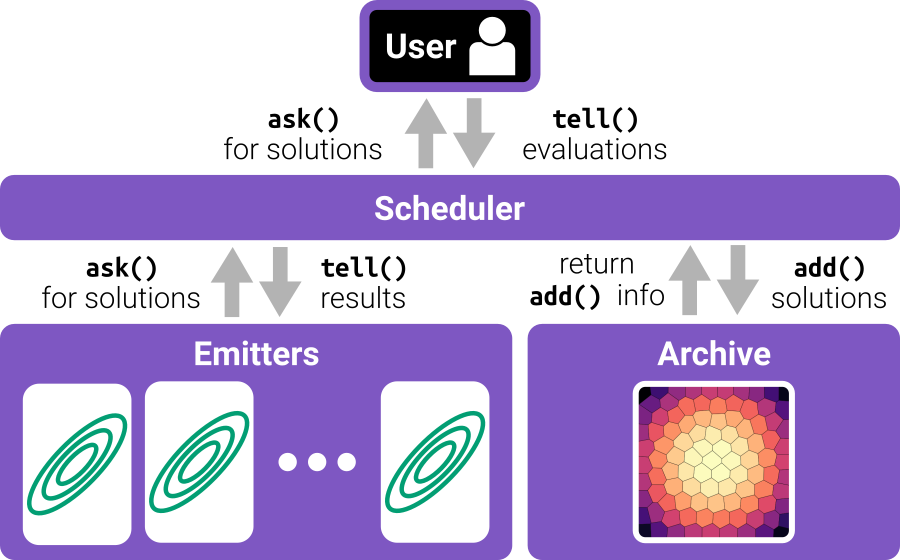
Following the RIBS framework (shown in the figure below), a standard algorithm in pyribs operates as follows:
The user calls the
ask()method on the scheduler. The scheduler requests solutions from each emitter by calling the emitter’sask()method.The user evaluates solutions to obtain the objective and measure values.
The user passes the evaluations to the scheduler’s
tell()method. The scheduler adds the solutions into the archive and receives feedback. The scheduler passes this feedback along with the evaluated solutions to each emitter’stell()method, and each emitter then updates its internal state.
Installation¶
pyribs supports Python 3.8 and above. The vast majority of users can install pyribs by running:
pip install ribs[visualize]
The command above includes the visualize extra. If you will not be using the pyribs visualization tools, you can install the base version of pyribs with:
pip install ribs
For more specific installation commands (e.g., installing from Conda or installing from source), visit the installation selector on our website.
To test your installation, import pyribs and print the version with this command:
python -c "import ribs; print(ribs.__version__)"
You should see a version number in the output.
Usage¶
Here we show an example application of CMA-ME in pyribs. To initialize the algorithm, we first create:
A 2D GridArchive where each dimension contains 20 cells across the range [-1, 1].
Three instances of EvolutionStrategyEmitter, all of which start from the search point 0 in 10-dimensional space and a Gaussian sampling distribution with standard deviation 0.1.
A Scheduler that combines the archive and emitters together.
After initializing the components, we optimize (pyribs maximizes) the negative 10-D Sphere function for 1000 iterations. Users of pycma will be familiar with the ask-tell interface (which pyribs adopted). First, the user must ask the scheduler for new candidate solutions. After evaluating the solution, they tell the scheduler the objectives and measures of each candidate solution. The algorithm then populates the archive and makes decisions on where to sample solutions next. Our toy example uses the first two parameters of the search space as measures.
import numpy as np
from ribs.archives import GridArchive
from ribs.emitters import EvolutionStrategyEmitter
from ribs.schedulers import Scheduler
archive = GridArchive(
solution_dim=10,
dims=[20, 20],
ranges=[(-1, 1), (-1, 1)],
)
emitters = [
EvolutionStrategyEmitter(
archive,
x0=[0.0] * 10,
sigma0=0.1,
) for _ in range(3)
]
scheduler = Scheduler(archive, emitters)
for itr in range(1000):
solutions = scheduler.ask()
# Optimize the 10D negative Sphere function.
objective_batch = -np.sum(np.square(solutions), axis=1)
# Measures: first 2 coordinates of each 10D solution.
measures_batch = solutions[:, :2]
scheduler.tell(objective_batch, measures_batch)
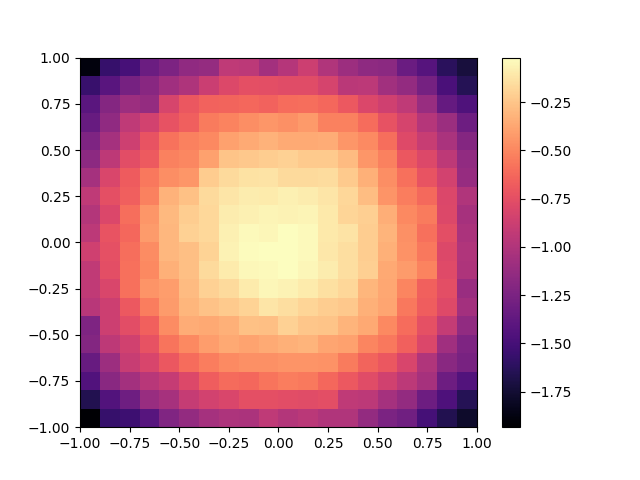
To visualize this archive with Matplotlib, we then use the grid_archive_heatmap function from ribs.visualize.
import matplotlib.pyplot as plt
from ribs.visualize import grid_archive_heatmap
grid_archive_heatmap(archive)
plt.show()
Documentation¶
The documentation is available online here. We suggest that new users start with the tutorials.
Paper and Citation¶
Two years after the initial release of pyribs, we released a paper that elaborates on the RIBS framework and the design decisions behind pyribs. For more information on this paper, see here. If you use pyribs in your research, please consider citing this paper as follows. Also consider citing any algorithms you use as shown below.
@inproceedings{10.1145/3583131.3590374,
author = {Tjanaka, Bryon and Fontaine, Matthew C and Lee, David H and Zhang, Yulun and Balam, Nivedit Reddy and Dennler, Nathaniel and Garlanka, Sujay S and Klapsis, Nikitas Dimitri and Nikolaidis, Stefanos},
title = {Pyribs: A Bare-Bones Python Library for Quality Diversity Optimization},
year = {2023},
isbn = {9798400701191},
publisher = {Association for Computing Machinery},
address = {New York, NY, USA},
url = {https://doi.org/10.1145/3583131.3590374},
doi = {10.1145/3583131.3590374},
abstract = {Recent years have seen a rise in the popularity of quality diversity (QD) optimization, a branch of optimization that seeks to find a collection of diverse, high-performing solutions to a given problem. To grow further, we believe the QD community faces two challenges: developing a framework to represent the field's growing array of algorithms, and implementing that framework in software that supports a range of researchers and practitioners. To address these challenges, we have developed pyribs, a library built on a highly modular conceptual QD framework. By replacing components in the conceptual framework, and hence in pyribs, users can compose algorithms from across the QD literature; equally important, they can identify unexplored algorithm variations. Furthermore, pyribs makes this framework simple, flexible, and accessible, with a user-friendly API supported by extensive documentation and tutorials. This paper overviews the creation of pyribs, focusing on the conceptual framework that it implements and the design principles that have guided the library's development. Pyribs is available at https://pyribs.org},
booktitle = {Proceedings of the Genetic and Evolutionary Computation Conference},
pages = {220–229},
numpages = {10},
keywords = {framework, quality diversity, software library},
location = {Lisbon, Portugal},
series = {GECCO '23}
}
Contributors¶
pyribs is developed and maintained by the ICAROS Lab at USC. For information on contributing to the repo, see CONTRIBUTING.
Nikitas Klapsis
We thank Amy K. Hoover and Julian Togelius for their contributions deriving the CMA-ME algorithm.
Users¶
pyribs users include:
Adaptive & Intelligent Robotics Lab (Imperial College London)
Various researchers at the University of Tsukuba
Publications¶
For the list of publications that use pyribs, refer to our Google Scholar entry.
Software¶
See the GitHub dependency graph for the public GitHub repositories which depend on pyribs.
Citing Algorithms in pyribs¶
If you use the following algorithms, please consider citing their relevant papers:
CMA-ME: Fontaine 2020
@inproceedings{10.1145/3377930.3390232, author = {Fontaine, Matthew C. and Togelius, Julian and Nikolaidis, Stefanos and Hoover, Amy K.}, title = {Covariance Matrix Adaptation for the Rapid Illumination of Behavior Space}, year = {2020}, isbn = {9781450371285}, publisher = {Association for Computing Machinery}, address = {New York, NY, USA}, url = {https://doi.org/10.1145/3377930.3390232}, doi = {10.1145/3377930.3390232}, booktitle = {Proceedings of the 2020 Genetic and Evolutionary Computation Conference}, pages = {94–102}, numpages = {9}, location = {Canc\'{u}n, Mexico}, series = {GECCO '20} }CMA-MEGA: Fontaine 2021
@inproceedings{NEURIPS2021_532923f1, author = {Fontaine, Matthew and Nikolaidis, Stefanos}, booktitle = {Advances in Neural Information Processing Systems}, editor = {M. Ranzato and A. Beygelzimer and Y. Dauphin and P.S. Liang and J. Wortman Vaughan}, pages = {10040--10052}, publisher = {Curran Associates, Inc.}, title = {Differentiable Quality Diversity}, url = {https://proceedings.neurips.cc/paper/2021/file/532923f11ac97d3e7cb0130315b067dc-Paper.pdf}, volume = {34}, year = {2021} }
CMA-MAE: Fontaine 2022
@misc{cmamae, doi = {10.48550/ARXIV.2205.10752}, url = {https://arxiv.org/abs/2205.10752}, author = {Fontaine, Matthew C. and Nikolaidis, Stefanos}, keywords = {Machine Learning (cs.LG), Artificial Intelligence (cs.AI), FOS: Computer and information sciences, FOS: Computer and information sciences}, title = {Covariance Matrix Adaptation MAP-Annealing}, publisher = {arXiv}, year = {2022}, copyright = {arXiv.org perpetual, non-exclusive license} }
Scalable CMA-MAE: Tjanaka 2022
@ARTICLE{10243102, author={Tjanaka, Bryon and Fontaine, Matthew C. and Lee, David H. and Kalkar, Aniruddha and Nikolaidis, Stefanos}, journal={IEEE Robotics and Automation Letters}, title={Training Diverse High-Dimensional Controllers by Scaling Covariance Matrix Adaptation MAP-Annealing}, year={2023}, volume={8}, number={10}, pages={6771-6778}, keywords={Covariance matrices;Training;Neural networks;Legged locomotion;Reinforcement learning;Evolutionary robotics;Evolutionary robotics;reinforcement learning}, doi={10.1109/LRA.2023.3313012} }
Additional QD Libraries¶
QDax: Implementations of QD algorithms in JAX. QDax is suitable if you want to run entire QD algorithms on hardware accelerators in a matter of minutes, and it is particularly useful if you need to interface with Brax environments.
qdpy: Python implementations of a wide variety of QD algorithms.
sferes: Contains C++ implementations of QD algorithms; can also handle discrete domains.
License¶
pyribs is released under the MIT License.
Credits¶
The pyribs package was initially created with Cookiecutter and the audreyr/cookiecutter-pypackage project template.



